RESEARCH
More than half of the ice-free land surface on Earth is directly modified by humans, and of this, over two-thirds is occupied by cropland and pastures [1]. To preserve remaining natural landscapes while meeting the needs of a growing human population, human-altered land must be optimized, not expanded. This requires enhancing the efficiency of agriculture, while rehabilitating lands that provide little functional value (e.g. contaminated sites, urban lawns) to states that contribute to ecological and human wellbeing.
|
Pushing the limits of soil function and plant productivity For most of the 20th century, traditional plant breeding and intensive chemical management led to substantial gains in crop production. In contrast, soil- and plant-associated microbiomes represent genetic and functional reservoirs that have barely been tapped. New high-throughput tools now allow us to ask questions about how microorganisms influence soil and plant function across agricultural and ecological landscapes, and how this might be altered. In our lab, we seek to understand and modify the factors that shape soil- and plant-associated microbiomes, while assessing their impact on important agricultural and ecological processes. |
|
Using high-throughput sequencing, custom-designed in-soil selection systems, and other approaches from molecular biology, ecology, soil science, and plant science, our lab aims to better understand and optimize soil- and plant-associated microbiomes. Projects typically address at least one of three main areas:
1 - Hooke and Martín-Duque. 2012. Land transformation by humans: a review. GSA Today 22: 4-10. |
|
Ongoing projects
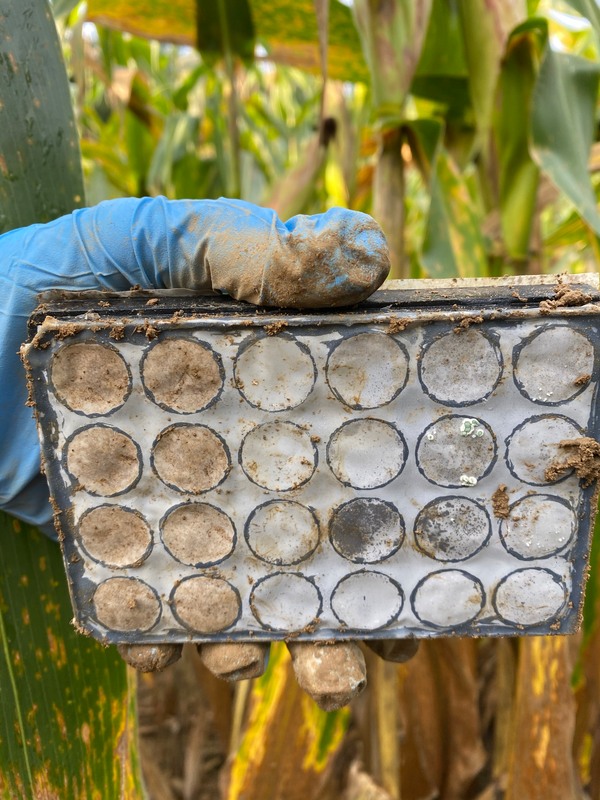
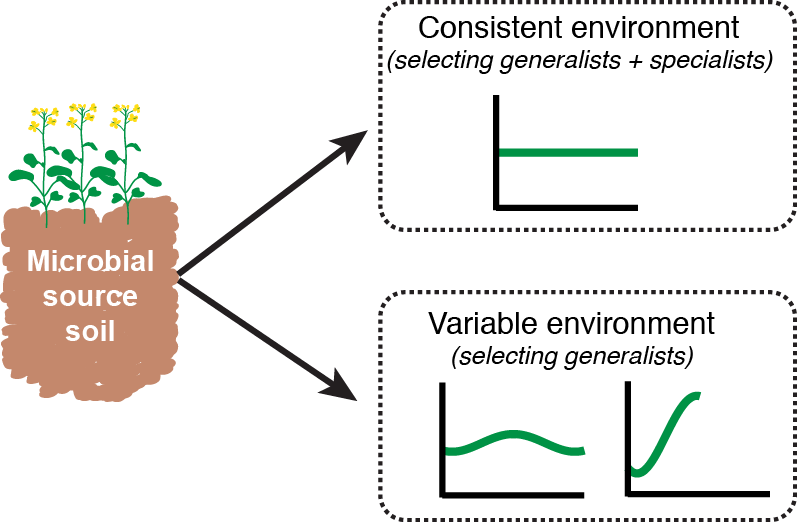
Specialist and generalist microorganisms in agricultural systems Funded by the USDA Foundational Program (PI is TH Bell) Agriculture typically reduces biodiversity relative to unmanaged environments, selecting against specialist organisms that survive under narrow conditions, while promoting generalists that tolerate a broader environmental range. Various management practices (e.g. nutrient applications) impose cyclical pressures that microorganisms must contend with, and that widen intra-annual variability in the environment. Shifting towards generalist-dominated communities could have landscape-scale consequences for key soil processes. In this project, we impose differing environmental gradients in custom in-soil selection systems, allowing us to collect generalists and specialists of different types. We then assess the taxonomy and function of these assemblages. Ultimately, we aim to augment functional and ecological annotations of sequence data, and better inform microbial management. |
|
Is there a role for microbial management in organic agriculture?
Funded by the USDA Organic Transitions Program (Lead PI is TH Bell; Co-PIs are Jason Kaye, Anna Busch) Many farmers believe soil microorganisms are key to soil functioning, but are unsure how to effectively manage microbial populations. Attempts to directly manage microorganisms are common, through approaches like compost development, Korean natural farming, and microbial product applications. However, the complexity and invisibility of microbial communities make it hard to benchmark potential solutions, leaving farmers without a framework for microbial management. In this project, we examine how soil type and farming practices interact with passive and active microbial management, in collaboration with farmer partners, and within a long-term organic project at Penn State. Specifically, we assess natural microbial colonization of farmed soils, the potential for forced microbial colonization of these soils, and the value of a widely used metric for assessing microbial health. We also interact with farmers through annual meetings, farmer conferences, and Extension publications. |
|
Interactions between soil microbiomes and soil surface consortia
Funded by the Joint Genome Institute (Lead PI is TH Bell; Co-PI is Mary Ann Bruns) We are assessing changes in the metatranscriptome of a Cyanobacteria-based soil surface consortium in the presence and absence of underlying soil microbiomes. This project aims to understand the degree to which in-soil competition can alter the function of microbial products for agriculture. |
In-soil persistence of genetically modified bacterial strains
(Lead PI is Howard Salis; Co-PI is TH Bell)
We are assessing how different soil factors and application approaches impact the in-soil persistence of bacteria with and without genetic modifications.
(Lead PI is Howard Salis; Co-PI is TH Bell)
We are assessing how different soil factors and application approaches impact the in-soil persistence of bacteria with and without genetic modifications.
|
Root-centric microbiome analysis
Funded by the USDA Foundational Program (Lead PI is Michela Centinari; Co-PIs are TH Bell, David Eissenstat) We are examining the influence of root order, age, depth, and spatial arrangement on microbial recruitment. Whereas most studies of root-associated microorganisms use homogenized root systems, we seek to understand the fine-scale factors that shape root-microbe interactions at different points in space. The primary work for this project takes place in an experimental vineyard at Rock Springs in Pennsylvania, using root visualization windows within 1 metre deep root observation boxes. This field setup is unique, and allows us to sample grapevine roots across a substantial depth profile. |
Resilient switchgrass cultivars for marginal lands
Funded by DOE/USDA (Lead PI is John Carlson; Co-PIs are TH Bell, Jesse Lasky, Marvin Hall, Stacy Bonos, Julie Hansen, Donald Viands)
We are assessing relationships between switchgrass genotypes, microbiome composition, and switchgrass performance in multiple marginal sites in the Northeast US.
Pennsylvania Soybean On-Farm Network
Funded by Pennsylvania Soybean Board (Lead PI is Paul Esker; Co-PIs are TH Bell, Delbert Voight, Elizabeth Bosak, Andrew Frankenfield, Heidi Reed, Alyssa Collins)
We are assessing the association of a variety of potentially yield-limiting factors, including microbial composition, with soybean growth across Pennsylvania.
Funded by DOE/USDA (Lead PI is John Carlson; Co-PIs are TH Bell, Jesse Lasky, Marvin Hall, Stacy Bonos, Julie Hansen, Donald Viands)
We are assessing relationships between switchgrass genotypes, microbiome composition, and switchgrass performance in multiple marginal sites in the Northeast US.
Pennsylvania Soybean On-Farm Network
Funded by Pennsylvania Soybean Board (Lead PI is Paul Esker; Co-PIs are TH Bell, Delbert Voight, Elizabeth Bosak, Andrew Frankenfield, Heidi Reed, Alyssa Collins)
We are assessing the association of a variety of potentially yield-limiting factors, including microbial composition, with soybean growth across Pennsylvania.
|
Previous projects
Selecting for desirable microbiome functions Funded by the Atkinson Center for a Sustainable Future This project focused on the potential of root-associated microbiome breeding to influence Brassica rapa biomass in salt-amended and untreated soils, using replicated selection lines. We have also sequenced root-associated microbiomes at each generation to look for links between microbiome composition and B. rapa growth, and to determine the predictability of targeted microbiome selection. |
|
Relationships between soil contaminants, willows, and soil microbiomes
Funded by Genome Canada and FQRNT I worked as a postdoctoral fellow on a large interdisciplinary project that aimed to enhance phytoremediation efficacy in willows. We used a variety of genomic approaches to examine the relationships between willows, microorganisms, soil characteristics, and contaminant uptake. These studies led us to understand that willow-microbe relationships vary in response to a number of factors, including willow species and soil contaminant concentrations. A better understanding of how plant-microbe relationships are influenced by the environment will allow us to characterize variability in the effectiveness of phytotechnologies. Ideally, this will eventually lead to more predictable outcomes when "green" technologies such as phytoremediation are applied, allowing them to better compete in the market with approaches that have a much larger environmental impact (e.g. dig and dump). |
|
Predictability of microbiome composition and function following contamination
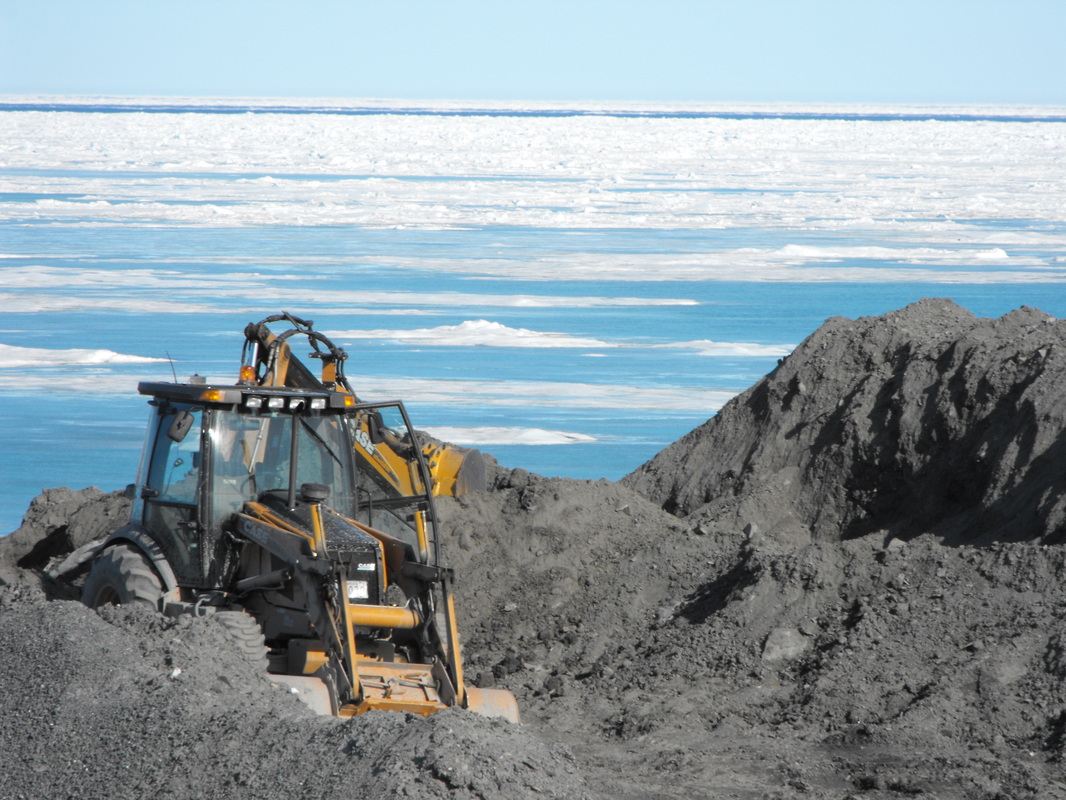
Funded by NSERC Many soil microbes can degrade petroleum, since they are adapted to metabolizing complex carbon compounds, such as those found in soil organic matter. This creates the potential for competition within the contaminated soil microbiome, just as we would be likely to find in most "pristine" microbial habitats. The focus of my Ph.D. work was the bioremediation of Arctic soils that have been contaminated by petroleum hydrocarbons. Nitrogen-based fertilizers are frequently added to hydrocarbon-contaminated soils, since they can allow increased microbial growth when carbon is in abundance and nitrogen is limiting, but not all hydrocarbon degraders are created equal. So which microorganisms benefit from this added nitrogen? What other factors might affect community composition in a contaminated soil? And are we promoting the most efficient hydrocarbon-degrading communities? |
|
Impacts of winter warming on soil microbial function
Our lab used a long-term field experiment to look at the effects of climate change on an old-field ecosystem. Infrared heaters warmed plots either year-round, or during the winter only, in an attempt to isolate the effects of climate change on winter processes. My role in this project was to examine how these factors affected microbial biomass, extracellular enzyme activity, and fungal:bacterial ratios. See here for more information about this project. |